VIDEO and FREE TRIAL
Qlucore Omics Explorer video
Qlucore Omics Explorer free trial

Find out more about the rich set of features below. The Qlucore Omics Explorer base module includes all key features used by scientists on a regular basis for multi-omics data analysis (including RNA-seq bulk and single cell). The NGS module adds a wide range of tools and methods for Genome browser centric analysis.
The features are the same for cloud and local installations.
At Qlucore we spend significant time and effort on testing and evaluating new functionality before adding it. These carefully thought-out features make the program very easy to use. Feedback from users shows that using Qlucore Omics Explorer adds value in many ways and two examples are:
Because Qlucore Omics Explorer is so easy to use, you don't need to be an expert in analysis to use it. The current user base spans a broad range of professions, from bioinformaticians, biologists, statisticians and chemists to medical doctors, students and professors.
The program is developed to support the user with fast, simple and visual analysis of measured data considering publicly available information such as gene ontologies, pathways and other system biology information to maximize the output of the analysis. The program is compatible with a very wide range of Omics data, for example:
You reach all key functionality with one or two mouse clicks and the results of your actions are always presented to you in real time by a visual update. The visual approach makes it easy to publish results as well as working in teams.
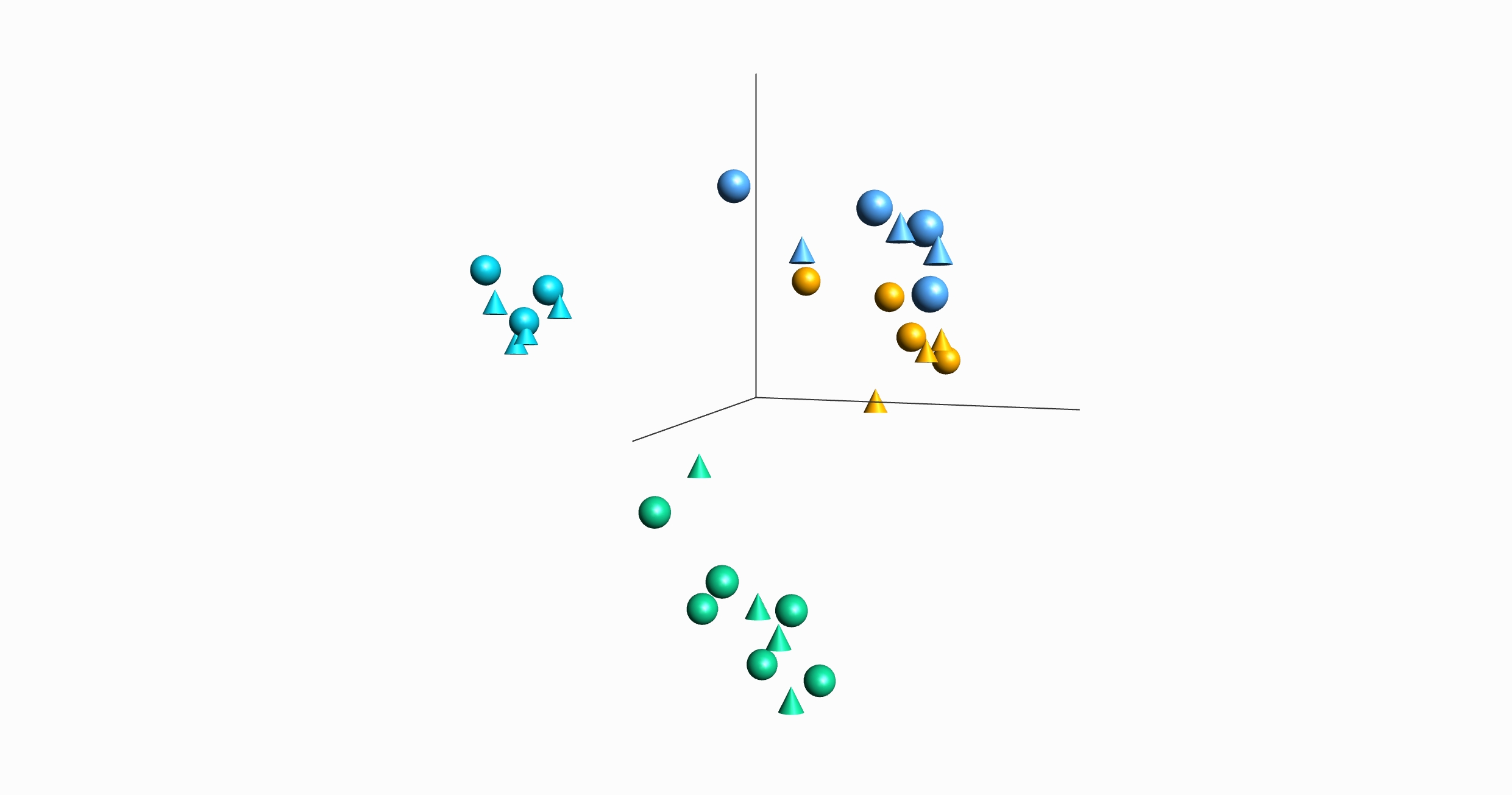
Qlucore Omics Explorer (QOE) ships in a base module with an option to add a NGS module with extensive functionality for NGS data analysis. Detailed information about the NGS module features is presented in the NGS Module feature overview document. Key features are highlighted below:
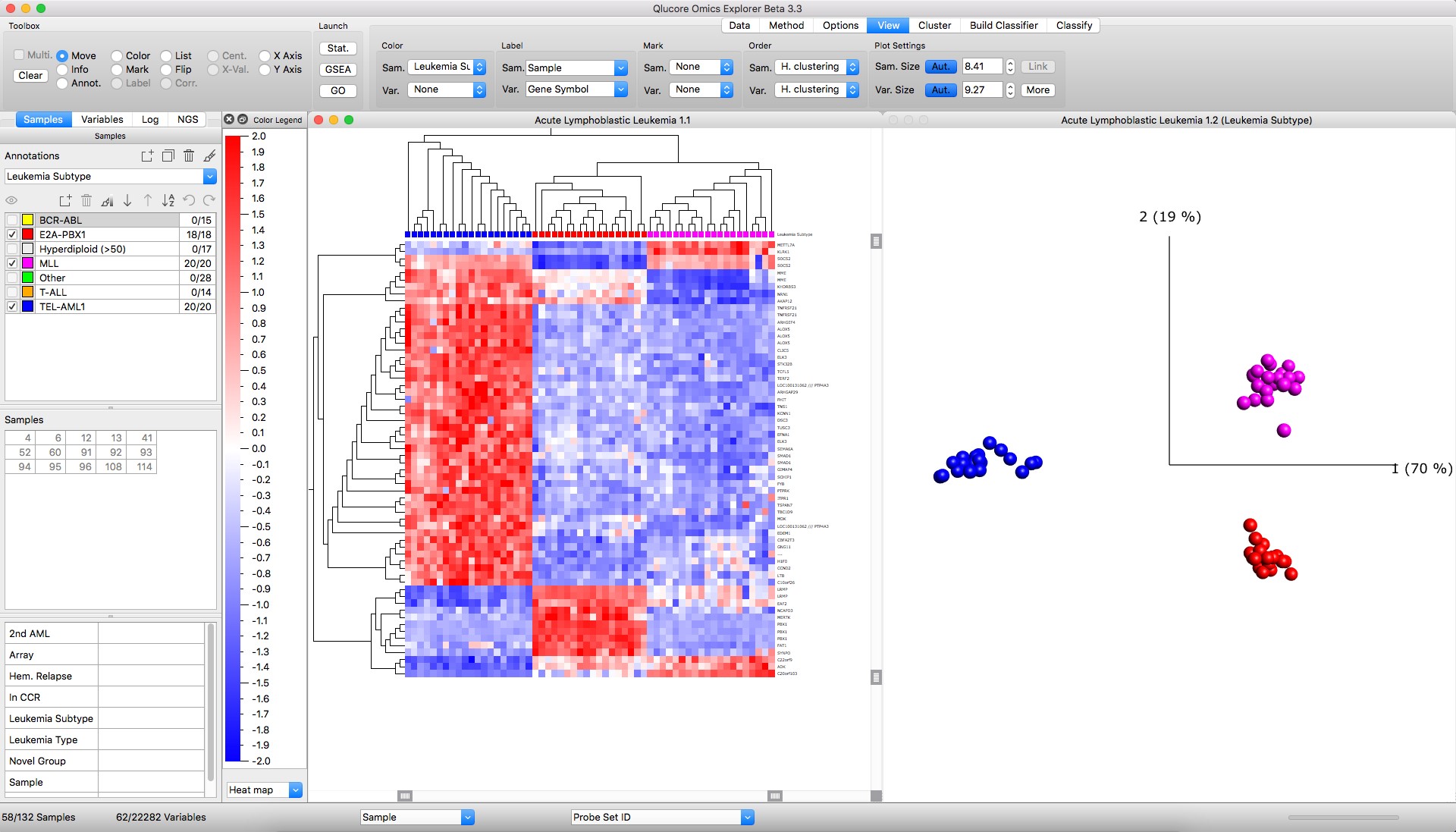
Figure: Left: Heatmap with hierarchical clustering Right: PCA sample plot
The Next Generation Sequencing (NGS) Module is an add-on module that will enable additional functionality related to analysis of data generated with NGS technologies and will make it possible to interactively and dynamically analyze and explore NGS data both from DNA and RNA.
All functionalities will be provided as integrated parts of Qlucore Omics Explorer and work with the well-known functionality.
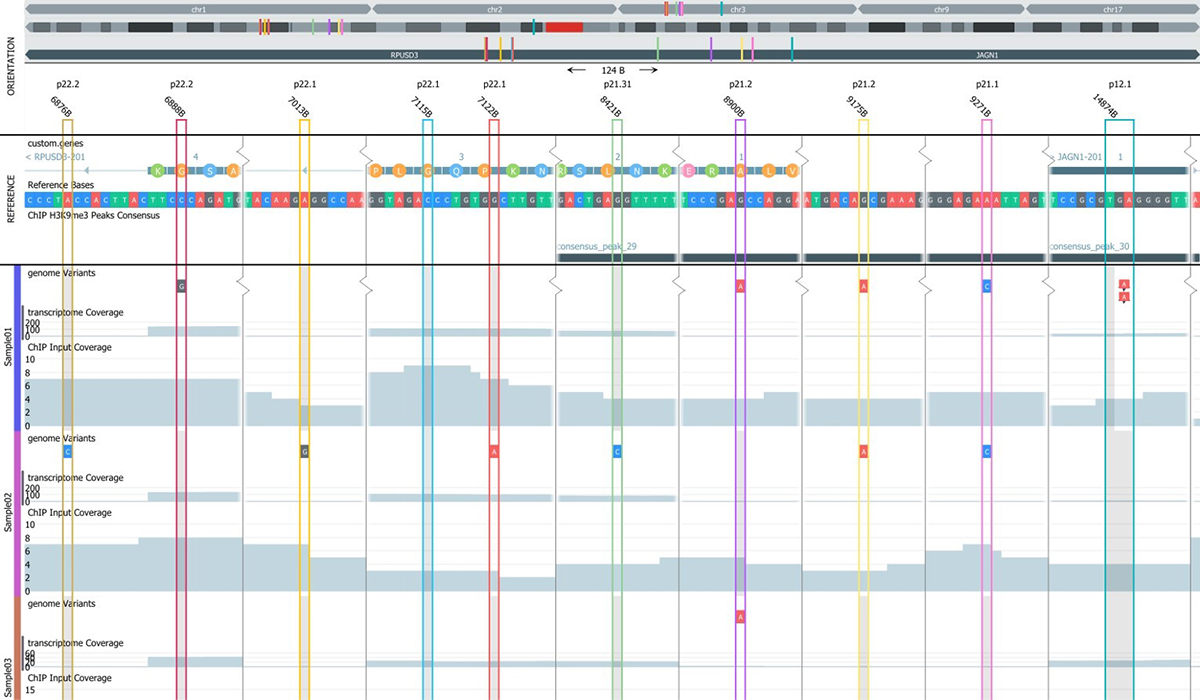
The main components of the NGS Module are:
The NGS module is unique in providing true interactive and dynamic analysis of NGS data. The Genome Browser content is dynamically updated when filters and filter cut-off are changed, for instance using sliders and check-boxes.
One unique feature of the NGS Module is the two-stage workflow. The first stage is a pre-processing step that prepares data for the fast and interactive analysis in stage two.
The project files and associated data for indexing samples is stored locally on the user’s computer to secure fast access and interactivity while files such as large BAM files can be stored anywhere on a network.
The options available for RNA-seq analysis are impressive. Utilizing the existing functionality in QOE for expression data and combining it with the new NGS functionality enables significantly increased analysis options. When a RNA-seq project is initiated, both quantitative normalized expression levels will be calculated at the same time as data is prepared for the genome browser. The program is built to seamlessly handle information both from a quantitative and genomic view. As an example, the user can define discriminating genes with a t-test, visualize them in a heat map and study the genes in the genome browser. If a sample group is selected or de-selected all plots will be updated and the
analysis can directly continue.
The peak analysis support in Qlucore Omics Explorer allows for comprehensive analysis of peak data such as ChIP-seq and ATAC-seq. The main components of the peak analysis processing are peak detection, consensus peaks and count matrix generation. The genome browser includes powerful filtering options for peak analysis, making it easy to find interesting features. The browser allows the user to annotate peaks and export the results as a bed file. If the experiment also includes RNA-seq data, it is possible to generate a count matrix for RNA-seq and swap between the RNA-seq and ChIP-seq count matrices during the analysis.
The Gene Fusion Workbench is shipped with the Mitelman reference data base. By combing filters of various types such as presence in the reference data base and quality, detection of both novel and well-established fusions is straightforward. The fusions can be thoroughly examined in the genome browser and visualized in the Circle plot for a great overview.
Read more about Next Generation Sequencing.
Qlucore Omics Explorer (QOE) is developed to enable fast, easy-to use and visual analysis of
measured data from a very wide range of sources and instruments transforming data into
insights considering publicly available information such as TCGA data, gene ontologies,
pathways and other system biology information to maximize the results and impact.
Qlucore Omics Explorer (QOE) supports the user with fast, simple and visual analysis of
measured data. The NGS Module is an add-on module that will enable additional functionality
related to data generated with NGS technologies and will make it possible to interactively and
dynamically analyze and explore NGS data both from DNA and RNA.